Understanding AMPylation and its Role in Neurons
Post-translational protein modifications play critical roles in a broad range of signaling pathways and biological processes in health and disease states. Modifications include the addition of specific chemical groups (e.g., phosphate, methyl, acetyl), polypeptides (e.g., ubiquitin, SUMO protein), and complex molecules (e.g., glycans, glycosylphosphatidylinositol, or GPI) with biologically significant outcomes. For example, protein phosphorylation is considered the most frequent post-translational modification and plays a central role in regulating enzymatic activity (Virag et al. 2020). Ubiquitination is critical in protein homeostasis through its ability to target proteins for proteasomal degradation. The addition of complex groups such as sugars or glycans is essential for protein localization and protein-protein interactions (Karve and Cheema, 2011, Virag et al. 2020). While the functional roles and biological outcomes of phosphorylation, glycation, and ubiquitination are well characterized, less is known about other post-translational modifications such as AMPylation.
What is AMPylation?
In AMPylation, specific protein residues are modified by the addition of an adenosine monophosphate (AMP) group. The AMPylation process relies on the enzymatic transfer of AMP from a donor ATP molecule to an available hydroxyl group in serine, threonine, or tyrosine. In humans, FIC (filamentation induced by cyclic AMP) domain containing (FICD/HYPE), and a recently identified pseudokinase, selenoprotein-O (SelO), are the only two known enzymes (ampylases) capable of catalyzing this transfer (Sreelatha et al. 2018, Chatterjee and Truttmann, 2021). FICD localizes to the ER, while SelO is found in the mitochondria, where these ampylases modify proteins implicated in various biological processes such as ER homeostasis and oxidative stress, respectively (Sanyal et al. 2020).
Developing new tools for detecting AMPylated proteins
Elucidating the biological functions of AMPylation is often challenging due to the lack of specific and sensitive antibodies to detect AMPylated protein residues. To overcome this challenge, Hopfner and colleagues partnered with GenScript. Leveraging on its expertise and strengths of its “Custom Monoclonal Antibody Generation” platform, they were able to successfully develop highly sensitive and specific monoclonal antibodies that detect AMPylation irrespective of its protein context (Hopfner et al. 2020).
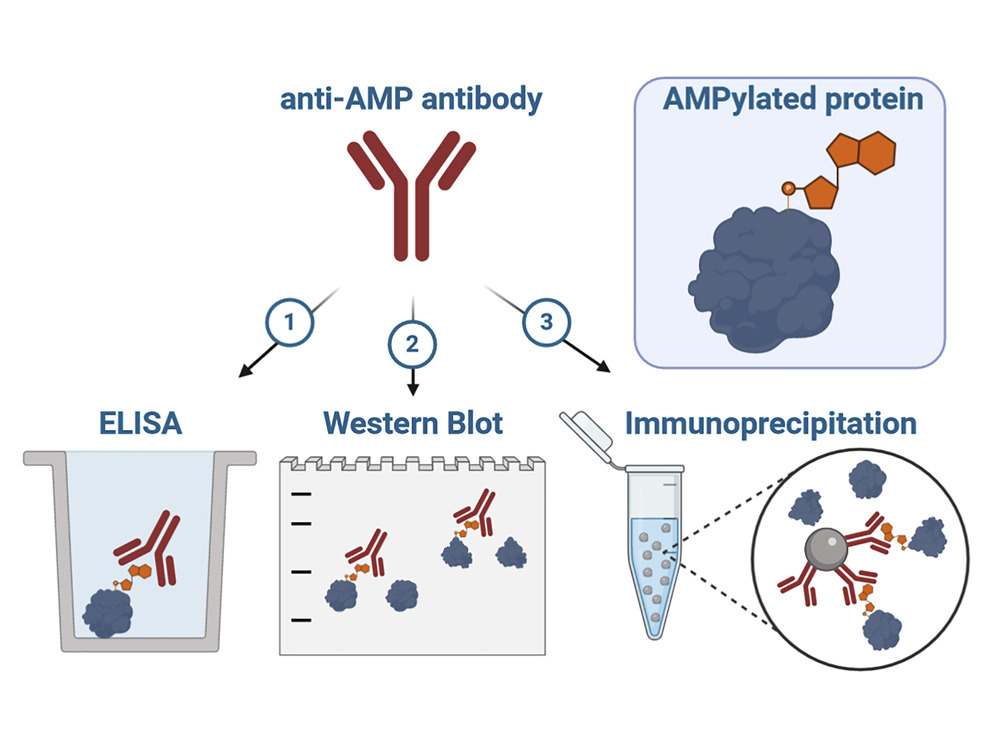
Graphical Abstract from Hopfner et al. 2021 ( https://creativecommons.org/licenses/by/4.0/)
Anti-AMP Mouse monoclonal antibody candidates were validated through ELISA and Western blotting to identify antibodies detecting AMP modifications irrespective of the protein backbone. Learn more about Hopfner’s strategy in Webinar-Development and Characterization of Monoclonal Antibodies Against AMPylation.
Roles of AMPylation in neurons
Several AMPylated protein targets have already been identified in neurons, suggesting that AMP-modification of proteins may serve different roles in the brain. For example, human FICD/HYPE binds and AMPylates alpha-synuclein, reducing its aggregation in vitro. Because FICD/HYPE is enriched in nigral dopaminergic neurons, Sanyal and colleagues have suggested that AMPylation may play a protective role in the Parkinson’s disease pathology (Sanyal et al. 2019).
Using a newly developed chemical-proteomic strategy, Kielkowski et al. 2020, identified several novel AMPylation targets in neurons, such as transport and cytoskeletal proteins. By modulating FICD activity in cerebral organoids, Kielkowski and colleagues demonstrated a role for AMPylation in neurogenesis and neuronal differentiation. Specifically, decreased FICD activity was associated with reduced neurogenesis, while its overexpression promoted neuronal differentiation.
More recently, the Kielkowski group applied the same chemical-proteomic strategy to uncover a propensity for increased AMPylation of lysosomal proteins in neuroblastoma cells (Kielkowski et al. 2020, Becker et al. 2021). Furthermore, by using iNGN cells (hiPSCs with inducible expression of Neurogenin-1 and -2) that rapidly differentiate into bipolar neurons, they showed that increased AMPylation of specific lysosomal targets (e.g., ACP2 and ABHD6) was associated with neuronal maturation.
AMPylation also occurs on PLD3, a lysosomal phospholipase recently implicated in Alzheimer’s disease. Using a combined Phos-tag/SDS-PAGE approach, the Kielkowski group confirmed increased PLD3 AMPylation in parallel with neuronal maturation (Nackenoff et al. 2021). Although the enzyme responsible for PLD3 AMPylation remains unidentified, these investigators discovered that AMPylated PLD3 had decreased catalytic activity, further raising questions about how AMPylation of PLD3 may be affected in Alzheimer’s disease.
Overall, recent studies have identified the presence of several AMPylated proteins in the brain, which merits further investigation to elucidate the exact role this post-translational modification plays in normal neuronal differentiation and disease states.
Reference
Becker, T. et al. AMPylation is a specific lysosomal protein posttranslational modification in neuronal maturation. bioRxiv (2021) 2021.03.02.433531; doi: https://doi.org/10.1101/2021.03.02.433531
Chatterjee, B. K. & Truttmann, M. C. Fic and non-Fic AMPylases: protein AMPylation in metazoans. Open Biol. (2021) doi:10.1098/rsob.210009.
Höpfner, D. et al. Monoclonal Anti-AMP Antibodies Are Sensitive and Valuable Tools for Detecting Patterns of AMPylation. iScience (2020) doi:10.1016/j.isci.2020.101800.
Karve, T. M. & Cheema, A. K. Small Changes Huge Impact: The Role of Protein Posttranslational Modifications in Cellular Homeostasis and Disease. J. Amino Acids (2011) doi:10.4061/2011/207691.
Kielkowski, P. et al. FICD activity and AMPylation remodelling modulate human neurogenesis. Nat. Commun. (2020) doi:10.1038/s41467-019-14235-6.
Nackenoff, A. G. et al. PLD3 is a neuronal lysosomal phospholipase D associated with β-amyloid plaques and cognitive function in Alzheimer’s disease. PLoS Genet. (2021) doi:10.1371/journal.pgen.1009406.
Sanyal, A. et al. Alpha-Synuclein Is a Target of Fic-Mediated Adenylylation/AMPylation: Possible Implications for Parkinson’s Disease. J. Mol. Biol. (2019) doi:10.1016/j.jmb.2019.04.026.
Sreelatha, A. et al. Protein AMPylation by an Evolutionarily Conserved Pseudokinase. Cell (2018) doi:10.1016/j.cell.2018.08.046.
Virág, D. et al. Current Trends in the Analysis of Post-translational Modifications. Chromatographia (2020) doi:10.1007/s10337-019-03796-9.
- Like (3)
- Reply
-
Share
About Us · User Accounts and Benefits · Privacy Policy · Management Center · FAQs
© 2026 MolecularCloud



