Neoantigen Peptides for Personalized T Cell Immunotherapies in Brain Cancer
Neoantigen Peptides in Personalized T Cell Immunotherapies for Medulloblastoma
What is medulloblastoma?
Medulloblastoma is a common malignant brain tumor in children, in fact, the most frequently diagnosed, accounting for over 60% of intracranial embryonal tumors. First described and histologically characterized in the 1920s, medulloblastomas start in the cerebellum and spread to other brain regions through the cerebrospinal fluid (CSF). Medulloblastoma results from sporadic mutations occurring in progenitor cells during embryonic brain development (Juraschka and Taylor 2019).
The heterogeneity of medulloblastoma has been a critical challenge to effective treatment. Based on histological findings, four main types of medulloblastomas are recognized, including classic (CLA), desmoplastic/nodular (DN), extensive nodularity (MBEN), and large cell/anaplastic (LCA). Unsurprisingly, together with their morphological heterogeneity, medulloblastoma tumors are also molecularly diverse. Four medulloblastoma molecular subgroups have been identified (i.e., WNT, SHH, Group3, and 4). These subgroups also differ in clinical presentation, with Group 3 and WNT types having the worst and best prognosis, respectively (Zou et al. 2020).
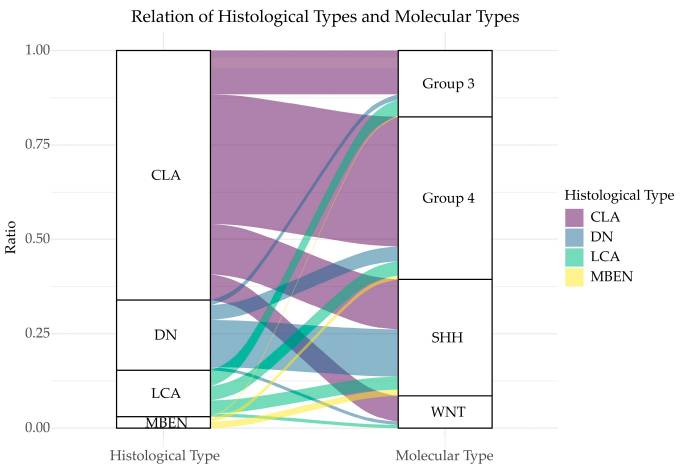
Medulloblastoma-Morphological and Molecular Groups. “Relation of histological types and molecular types. The two columns represent histological classification and molecular classification, respectively. Different heights correspond to different ratios. Lines between the columns represent the overlapping classification systems. The broader a line, the more overlapping patients it has.” Retrieved from Figure 1 of Zou et al. 2020. Only a portion of the legend is included. https://creativecommons.org/licenses/by/4.0/
Therapies for medulloblastoma
The current therapy strategy for medulloblastoma combines tumor resection, craniospinal radiotherapy, and chemotherapy. The exact course of treatment is highly patient-specific and depends on the medulloblastoma subgroup and overall clinical presentation. While efficacious, leading to improved survival for even high-risk patients, the current multimodal therapy has adverse consequences, including neurological deficits, impaired cognition, and a host of other side effects (Juraschka and Taylor 2019). Additionally, disease relapse after treatment occurs in up to 33% of patients and carries a very low survival rate (Kumar et al. 2021).
Personalized immunotherapies for medulloblastoma
Recognizing the need for personalized therapies to tackle this highly heterogeneous disease, a team at George Washington University Cancer Center, Washington, leveraged a combination of genomic and proteomic approaches to identify neoantigen peptides in brain tumors from pediatric patients (Rivero-Hinojosa et al. 2021). Identifying tumor-specific antigens or neoantigens would support developing T cell cancer immunotherapies, which are expected to offer a more targeted and safer treatment for medulloblastoma patients.
With this goal in mind, Rivero-Hinojosa and colleagues developed a proteogenomic approach requiring low amounts of tumor tissue, as low as 10 mg, for tumor-specific antigen identification. This approach helps overcome limitations to neoantigen discovery imposed by the reduced mutation burden and low tissue availability typical of pediatric brain tumors.
For their proteogenomic workflow, Rivero-Hinojosa and colleagues first analyzed medulloblastoma tumor tissue samples by whole genome sequencing (WGS) and RNA-seq, which enabled them to identify several genomic events, including gene fusions, insertion/deletions (indels), single nucleotide variations, and abnormal splicing. The resulting tumor genomic database and proteomics data, enabled by LC-MS/MS analysis, supported the identification of peptides validated as tumor-specific by comparison against normal tissue samples and databases.
By following this workflow, investigators identified over 300 tumor-specific peptides from 46 medulloblastoma tumor samples, predominantly originating from abnormal splicing events. Therefore, their proteogenomic approach successfully identified multiple neoantigen candidates in each tumor type, which could be targeted with T cell immunotherapeutics.
Synthetic peptides support T cell therapeutic development
To test the clinical practicality of their approach, investigators used synthetic peptides having sequences matching that of neoantigens identified from a patient’s tumor. Autologous dendritic cells loaded with multiple tumor-specific peptides (~10 to 25 mer) synthesized by GenScript, were used for repeated stimulation of patient-derived peripheral blood mononuclear cells (PBMCs). The resulting expanded T cell population was characterized for activity and phenotype by assaying for interferon-gamma and tumor necrosis factor-alpha and membrane markers, such as CD3, CD8, and CD4. Rivero-Hinojosa and colleagues found that the expanded autologous T cells were equally comprised of CD4+ and CD8+ cells and primarily consisted of T effector memory (TEM) and T central memory (TCM) cell subsets.
Unfortunately, the team did not have at their disposal patient-derived tumor cell lines to demonstrate the efficacy of the tumor-specific T cell product generated. Therefore, Rivero-Hinojosa and colleagues applied their proteogenomic workflow to identify neoantigen peptides from medulloblastoma cell lines. Once peptides were identified and validated, the team was able to generate a T cell product that demonstrated significant cytolytic activity against corresponding medulloblastoma cells.
Overall, the proteogenomic workflow developed by Rivero-Hinojosa and colleagues provides new opportunities for generating highly specific personalized T cell immunotherapies for medulloblastoma based on the use of limited brain tumor samples. A personalized T cell immunotherapy targeting multiple tumor-specific antigens would help address the challenges of the high tumor heterogeneity typical in medulloblastoma tumors. Additionally, highly specific T cell products have the potential to improve upon the toxicity associated with current treatments.
How much do you know about Medulloblastoma?
Take the Quiz, test your brain tumor knowledge, and try your luck at some cool prizes.
- Like (3)
- Reply
-
Share
About Us · User Accounts and Benefits · Privacy Policy · Management Center · FAQs
© 2026 MolecularCloud



