Leveraging CRISPR/Cas9 to target fusion oncogenes in vivo for selective elimination of cancer cells
Fusion oncogenes are
strong driven factor for tumor progress, which only express in cancer cell. Researchers
observed that eliminating directionally fusion oncogenes where the cancer induced
by that, giving rise to apoptosis of cancer cells. Though this therapeutic
target is very potential, directional killing specific cell is more challenging.
For conquering the critical problem, the researchers devised a simple,
efficient, and non-patient-specific gene-editing strategy, based on CRISPR/Cas9
technology, that is able to specifically destroy fused oncogenes in cancer
cells by targeting the two introns involved in the rearrangement. The study was published in Nature Communications with the
heading: "In Vivo CRISPR/Cas9 targeting of Fusion Oncogenes for
Selective Elimination of Cancer Cells [1]."
What is fusion oncogene?
Fusion gene refers to a
chimera that connects the coding regions of two or more genes end to end and
puts them under the control of the same set of regulatory sequences including
promoters, enhancers, ribosome binding sequences, terminator, etc. That is an abnormal
result of incorrect connection of DNA fragment from two different genes, the accidental circumstance during cell division. Fusion
gene and its encoding protein, like transcription factor or tyrosine kinase, will
trigger cell uncontrolled proliferation and tumorigenesis. At present, people
have found that fusion genes are widely present in nearly 20% of cancer types
such as prostate tumor, breast cancer, lung tumor, and brain
tumor.
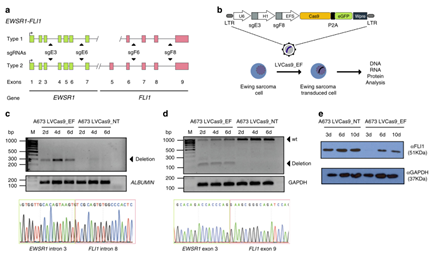
Selective elimination of
tumor cells by CRISPR/Cas9
After a series of
attempts, researchers have developed a targeted fusion oncogene editing method
based on CRISPR/Cas9 technology, and conducted on the EWSR1-FLI1 fusion gene.
This method targets two intron sequences of fusion oncogenes, which can induce
cancer cell-specific genome deletions, thereby eliminating key protein domains or
changing the reading frame of the fusion oncogene. It is worth noting that this
CRISPR/Cas9-based method only induces deletions in cells containing fusion
oncogenes, but does not affect the exon sequence or the protein expression of
the germline non-rearranged alleles. The same two guide RNA (sgRNAs) allow
specific breakpoints of different subtypes of or
specific fusion oncogenes for each patient, so it is a versatile method for treating
cancer-related fusion oncogenes.
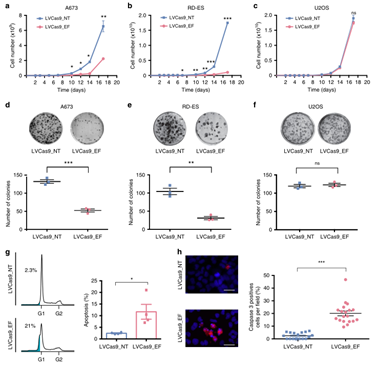
Reference:
[1]
Martinez-Lage, M et al. “In vivo CRISPR/Cas9 targeting of fusion
oncogenes for selective elimination of cancer cells.” Nature
communications vol. 11,1 5060. 8 Oct. 2020,
doi:10.1038/s41467-020-18875-x
- Like (1)
- Reply
-
Share
About Us · User Accounts and Benefits · Privacy Policy · Management Center · FAQs
© 2026 MolecularCloud




