Overview of Gibson Assembly®
Gibson Assembly® is a recombination-based molecular cloning method for the in vitro assembly of DNA fragments. Developed by Daniel G. Gibson and his colleagues in 2009, this methodology enables easy assembly of multiple DNA fragments into a circular plasmid in a single-tube isothermal reaction. The result is a scarless DNA molecule of up to 15 kb in size.
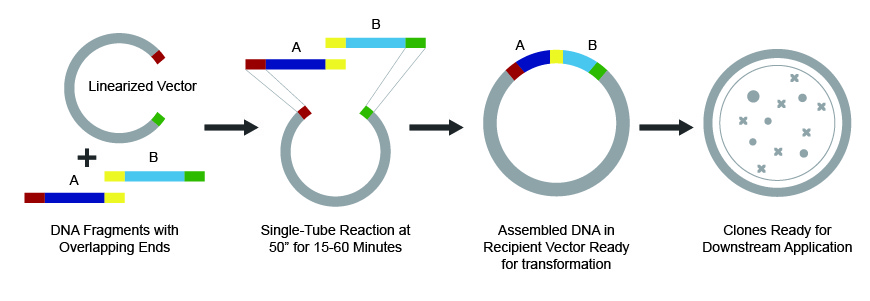
In this method, first fragments with 15-80 bp overlap with adjacent DNA parts are designed and synthesized. After synthesis, fragments are mixed with a reaction buffer containing three different types of enzyme: a T5 exonuclease, a DNA polymerase and a DNA ligase.
- The T5 exonuclease chews back both strands of DNA fragments from the 5’ end, generating 3’ single-stranded overhangs that can anneal with adjacent DNA fragments;
- The DNA polymerase then incorporates nucleotides to fill in the gaps within the annealed fragments;
- The DNA Ligase finally seals nicks in the assembled DNA to generate a double-stranded, fully-sealed DNA molecule.
Application and Advantages of Gibson Assembly®
Gibson Assembly can be used for seamless assembly of multiple fragments to generate natural or de novo DNA molecules as well as construction of DNA libraries for diverse down-stream applications. It can also be used for site-directed mutagenesis such as insertions, deletions and point mutations.
Compared with the conventional restriction enzyme/ligation-based cloning method, Gibson Assembly® offers many advantages:
- The assembly efficiency is significantly higher than the restriction enzyme/ligation-based cloning method;
- Greater number of fragments can be simultaneously combined in a single-tube reaction;
- There won’t be any concern for internal restriction enzyme cutting within a particular sequence;
- The procedure is faster and more efficient since it doesn’t require digestion and isolation of DNA fragments.
