2019 Highlights: What are the breakthroughs published in Science?
At the beginning of 2020, we look back to 2019 and sort out a collection of groundbreaking research published in Cell, Nature, and Science, hoping that the articles can help you stay connected with the latest research in molecular biology, gene editing technology, cell therapy, and immunology.
[1] Hachimoji DNA and RNA: A genetic system with eight building blocks
Highlights: The study reports DNA- and RNA-like systems built from eight nucleotide “letters” (hence the name “hachimoji”) that form four orthogonal pairs, makes available a mutable genetic system built from eight different building blocks. With increased information density over standard DNA and predictable duplex stability, hachimoji DNA has great potential applications. In addition to its technical applications, this work expands the scope of the structures that might support life in the cosmos.
In this study, the researchers report DNA- and RNA-like systems built from eight nucleotide “letters” (hence the name “hachimoji”) that form four orthogonal pairs. These synthetic systems meet the structural requirements needed to support Darwinian evolution, including a polyelectrolyte backbone, predictable thermodynamic stability, and stereoregular building blocks that fit a Schrödinger aperiodic crystal. Measured thermodynamic parameters predict the stability of hachimoji duplexes, allowing hachimoji DNA to increase the information density of natural terran DNA. Three crystal structures show that the synthetic building blocks do not perturb the aperiodic crystal seen in the DNA double helix. Hachimoji DNA was then transcribed to give hachimoji RNA in the form of a functioning fluorescent hachimoji aptamer. These results expand the scope of molecular structures that might support life, including life throughout the cosmos.
Reference: Hoshika, Shuichi, et al. "Hachimoji DNA and RNA: A genetic system with eight building blocks." Science 363.6429 (2019): 884-887.
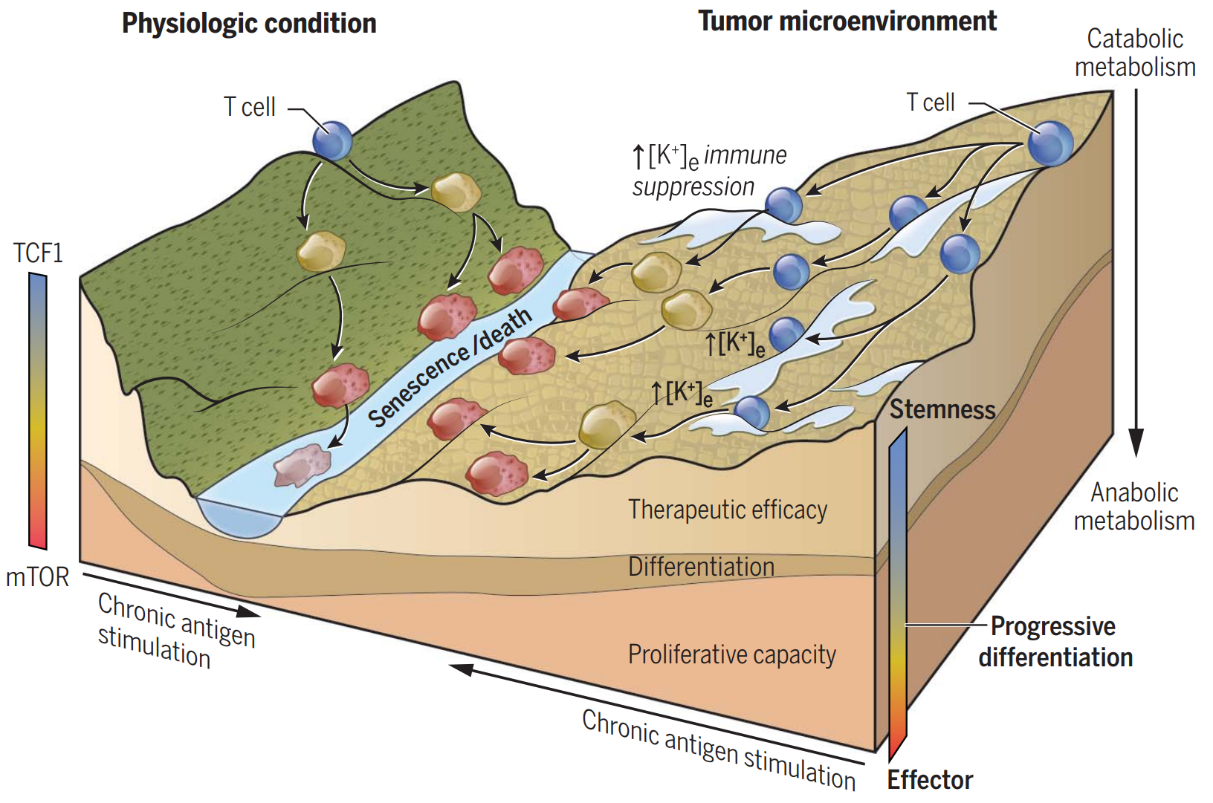
[2] T cell stemness and dysfunction in tumors are triggered by a common mechanism
Highlights: The findings deepen our understanding of how cancer can progress despite the presence of T cells that continue to harbor the capacity for its destruction, and finally identify new therapeutic strategies to metabolically induce stemness programs in antitumor T cells that enhance cancer immunotherapies.
In this study, the researcher found that the overabundance of potassium in the tumor microenvironment trigger suppression of T cell effector function while preserve stemness. High levels of extracellular potassium constrain T cell effector programs by limiting nutrient uptake, thereby inducing autophagy and reduction of histone acetylation at effector and exhaustion loci, which in turn produces CD8+ T cells with improved in vivo persistence, multipotency, and tumor clearance. This mechanistic knowledge advances our understanding of T cell dysfunction and may lead to novel approaches that enable the development of enhanced T cell strategies for cancer immunotherapy.
Reference: Vodnala, Suman Kumar, et al. "T cell stemness and dysfunction in tumors are triggered by a common mechanism." Science 363.6434 (2019): eaau0135.
[3] Reactivation of PTEN tumor suppressor for cancer treatment through inhibition of a MYC-WWP1 inhibitory pathway
Highlights: This study identified the MYCWWP1 axis as a fundamental and evolutionary conserved regulatory pathway for PTEN and PI3K signaling. This pathway emerges not only as a rheostat for growth control in physiological conditions but also as a critical vulnerability hijacked for neoplastic transformation, which may be reversed by WWP1 pharmacological inactivation. These findings pave the way toward a long-sought tumor suppressor “reactivation” approach to cancer treatment.
PTEN is a critical tumor suppressive phosphatase that is active in its dimer configuration at the plasma membrane. Polyubiquitination by the ubiquitin E3 ligase WWP1 (WW domain–containing ubiquitin E3 ligase 1) suppressed the dimerization, membrane recruitment, and function of PTEN. Either genetic ablation or pharmacological inhibition of WWP1 triggered PTEN reactivation and unleashed tumor suppressive activity. WWP1 appears to be a direct MYC (MYC proto-oncogene) target gene and was critical for MYC-driven tumorigenesis. This study identified indole-3-carbinol, a compound found in cruciferous vegetables, as a natural and potent WWP1 inhibitor. Thus, these findings unravel a potential therapeutic strategy for cancer prevention and treatment through PTEN reactivation.
Reference: Lee, Yu-Ru, et al. "Reactivation of PTEN tumor suppressor for cancer treatment through inhibition of a MYC-WWP1 inhibitory pathway." Science 364.6441 (2019): eaau0159.

